The Human Holobiont
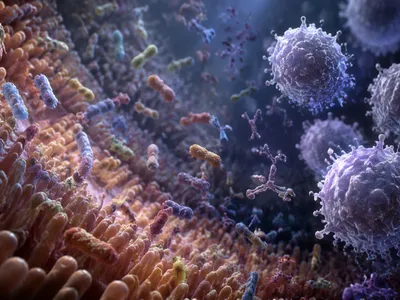
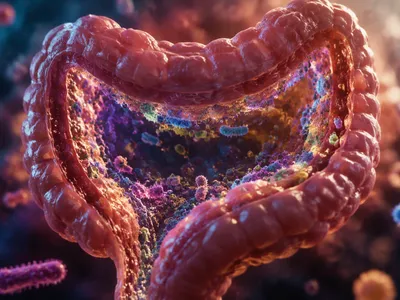
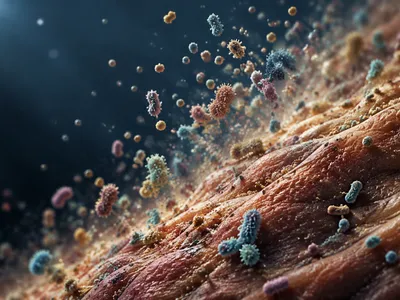
The contemporary immunological paradigm reconceptualizes the human organism not as a singular entity but as a holobiont—a superorganism comprising human cells and a vast consortium of commensal microorganisms. This shifts the focus from host-centric defense to a framework of integrated coexistence. The microbiota is now understood as a fundamental component of systemic immunity.
Evolutionary pressures have fostered a symbiotic relationship where microbial colonization is essential for calibrating immune reactivity. The constitutive presence of these communities provides continuous, low-level antigenic stimulation, priming the immune system for appropriate responses. This dialogue begins at birth and is influenced by factors such as delivery mode, diet, and antibiotic exposure. Disruption of this early colonization can have long-lasting immunological consequences.
Deciphering the Microbiota-Immune Dialogue
The crosstalk between host and microbiota occurs through multiple sophisticated channels. Pattern recognition receptors on epithelial and immune cells constantly sample microbial-associated molecular patterns. This surveillance is not merely for pathogn detection but for maintaining homeostatic tone.
Key immune adaptations include the priming of antimicrobial peptide production and the regulation of tight junction protein expression in the intestinal epithelium. Furthermore, dendritic cells extend dendrites into the gut lumen, sampling commensal antigens to orchestrate tolerogenic T-cell responses.
| Microbial Metabolite | Primary Immune Cell Target | Immunological Effect |
|---|---|---|
| Short-Chain Fatty Acids (e.g., Butyrate) | Regulatory T-cells (Tregs) | Promotes differentiation and function, enhancing peripheral tolerance. |
| Secondary Bile Acids | Natural Killer T (NKT) Cells | Modulates cytokine profiles, suppressing excessive inflammation. |
| Tryptophan Metabolites (e.g., Indole) | Aryl Hydrocarbon Receptor (AhR) in ILC3s | Stimulates IL-22 production, fortifying epithelial barrier integrity. |
This dynamic equilibrium ensures that commensals are contained within the mucosal lumen without eliciting destructive inflammatory responses. The failure of these regulatory mechanisms is a cornerstone in the pathogenesis of chronic inflammatory diseases.
Specific bacterial taxa, like segmented filamentous bacteria, are potent inducers of Th17 cells in the lamina propria, demonstrating how discrete organisms can shape the local immune landscape. This level of specificity underscores the precision of host-microbe interactions.
- Constant MAMP-PRR engagement maintains baseline immunocompetence, a state termed "tonic stimulation".
- Gut-associated lymphoid tissue development is structurally and functionally dependent on microbial signals.
- Microbiota influences the maturation and function of myeloid cells, including macrophages and neutrophils, in systemic tissues.
Microbial Molecules as Immune Instructors
Commensal microorganisms exert profound immunomodulatory effects primarily through the secretion of a diverse array of bioactive metabolites. These molecules serve as key instructors, shaping the development, function, and plasticity of the host immune system far beyond the gut mucosa.
Short-chain fatty acids (SCFAs) like butyrate, propionate, and acetate, produced by the fermentation of dietary fiber, are archetypal examples. Butyrate acts as a histone deacetylase inhibitor, epigenetically modulating gene expression in colonic Tregs and macrophages, thereby amplifying anti-inflammatory pathways and enhancing mucosal integrity.
Beyond SCFAs, other metabolites such as polysaccharide A from *Bacteroides fragilis* induce a shift towards regulatory T-cell responses, demonstrating bacterial strain-specific immunomodulation. Similarly, tryptophan derivatives engage the aryl hydrocarbn receptor in innate lymphoid cells, promoting interleukin-22 secretion and epithelial repair. The discovery that microbial products can mimic human signaling molecules, a phenomenon known as molecular mimicry, adds another layer of complexity to this cross-kingdom dialogue.
- Bile acid metabolism by gut bacteria generates secondary bile acids that act as signaling molecules via the FXR and TGR5 receptors, modulating inflammatory tone.
- Microbial-derived N-acyl amides can directly engage G-protein-coupled receptors on enteroendocrine and immune cells, influencing systemic metabolism and inflammation.
- Bacterial peptidoglycan fragments can translocate systemically, priming the innate immune system in the bone marrow and peripheral tissues, a concept known as "trained innate immunity."
This metabolic instruction is not a one-way street; host diet and genetics shape the microbial community and thus the metabolite landscape, creating a dynamic feedback loop that continuously fine-tunes immune homeostasis. Disruption of this metabolite production, termed metabolomic dysbiosis, is increasingly implicated in the etiology of autoimmune and metabolic disorders.
From Correlation to Mechanism
Early microbiota research established compelling correlations between microbial dysbiosis and disease states. The current frontier is definitively establishing causal mechanisms. This requires moving beyond sequencing surveys to experimental manipulation of specific microbial communities and their products in controlled models.
Gnotobiotic animal models, reared in sterile conditions and colonized with defined microbial consortia, have been instrumental. These studies allow researchers to dissect the immunological impact of introducing or removing specific bacterial species, directly linking microbial presence to immune phenotypes and providing proof of principle for causal relationships.
Advanced techniques like metagenomic sequencing coupled with metabolomics and transcriptomics enable a systems-level understanding. By integrating data on which genes are present, which metabolites are produced, and how host tissues respond, researchers can construct predictive models of host-microbe interactions. The integration of machine learning is now helping to decode these complex datasets and identify keystone species and functional pathways that are non-negotiable for immune health.
Fecal Microbiota Transplantation
The most direct clinical application of microbiota science is fecal microbiota transplantation, a procedure that epitomizes the shift from correlation to therapeutic intervention. By transferring a processed fecal suspension from a healthy donor to a recipient, FMT aims to restore a functional microbial ecosystem.
Its remarkable efficacy in treating recurrent *Clostridioides difficile* infection, which exceeds 90% in some studies, provides compelling proof-of-concept that manipulating the microbiome can resolve a severe pathological state. This success is attributed to the reintroduction of keystone species and functional networks that outcompete the pathogen and restore colonization resistance.
| Condition | FMT Status | Key Mechanistic Insight |
|---|---|---|
| Recurrent *C. difficile* | Standard of Care (in guidelines) | Restoration of bile acid metabolism and direct competition. |
| Ulcerative Colitis | Investigational (variable results) | Induction of mucosal healing via increased SCFA producers and regulatory immune cells. |
| Metabolic Syndrome | Early-phase trials | Transient improvement in insulin sensitivity linked to donor microbial engraftment. |
Beyond CDI, FMT is under rigorous investigation for conditions like inflammatory bowel disease, metabolic syndrome, and even neurological disorders. The variable outcomes in these trials highlight the complexity of the ecosystem being transferred; success may depend on precise donor-recipient matching, the recipient's baseline immune and metabolc status, and the engraftment of specific, yet often unidentified, bacterial consortia. Research is now focused on moving from whole-stool transplants to defined microbial consortia or "microbial pills" that can deliver a standardized, safe, and efficacious product, thereby eliminating the risks associated with undefined donor material.
FMT trials serve as unparalleled human discovery platforms. By analyzing responders versus non-responders, scientists can identify microbial signatures and metabolites that are causal to clinical improvement, thereby reverse-engineering the mechanisms of a successful transplant to inform next-generation probiotics and targeted therapies.