The Microscopic Architects of Our Gut
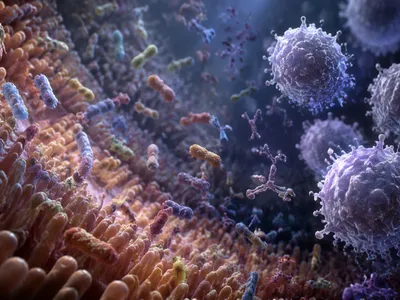
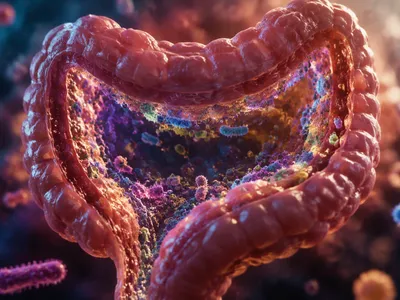
The human gastrointestinal tract harbors a complex ecosystem of microorganisms, collectively known as the gut microbiota, which functions as a metabolic organ influencing systemic health. Recent metagenomic analyses have revealed its profound role in immune modulation, nutrient absorption, and pathogen exclusion.
Dysbiosis, an imbalance in this community, is causally linked to numerous conditions far beyond digestive health. These include metabolic syndromes like obesity and type 2 diabetes, neuropsychiatric disorders, and autoimmune diseases, illustrating its systemic reach.
Central to this interaction is the gut-brain axis, a bidirectional communication network involving neural, endocrine, and immune pathways. Microbial metabolites, such as short-chain fatty acids (SCFAs) like butyrate, are key signaling molecules. Butyrate not only serves as the primary energy source for colonocytes but also exerts epigenetic influence by inhibiting histone deacetylases, thereby regulating gene expression in host cells and modulating inflammatory responses. Furthermore, the microbiota is essential for the development and function of the gut-associated lymphoid tissue (GALT), training the immune system to distinguish between commensals and pathogens, a process critical for maintaining immune homeostasis and preventing inappropriate inflammation.
- Production of essential vitamins (e.g., Vitamin K, B12) and bioactive compounds.
- Fermentation of non-digestible dietary fibers into beneficial short-chain fatty acids.
- Direct competition with pathogens for nutrients and adhesion sites (colonization resistance).
- Modulation of the intestinal barrier integrity and mucus layer composition.
Phage Therapy and the War Against Superbugs
The relentless rise of antimicrobial resistance (AMR) necessitates innovative therapeutic strategies, with bacteriophage therapy re-emerging as a promising precision antimicrobial tool. Phages are viruses that specifically infect and lyse bacterial cells, offering a targeted approach absent in broad-spectrum antibiotics.
A key advantage is their self-replicating nature at the site of infection and their ability to co-evolve with resistant bacteria. This dynamic interaction can potentially outmaneuver bacterial defense mechanisms, a significant limitationn of static antibiotic molecules.
Clinical applications have progressed from compassionate use to structured trials, particularly for treating drug-resistant Pseudomonas aeruginosa, Staphylococcus aureus, and Mycobacterium abscessus infections. Case studies involving prosthetic joint infections and chronic wound sepsis demonstrate significant biofilm disruption and clinical improvement where antibiotics had failed.
| Phage Type | Primary Target | Therapeutic Advantage |
|---|---|---|
| Lytic (Virulent) Phages | Active infection sites | Direct bacterial lysis; immediate reduction in pathogen load. |
| Lysogenic (Temperate) Phages | Not typically used directly | Can be engineered for gene delivery or as diagnostic tools. |
| Phage Cocktails | Poly-microbial or high-resistance-risk infections | Broadens spectrum; reduces chance of phage-resistant bacterial escape mutants. |
Despite promise, challenges include stringent phage purification to remove bacterial endotoxins, narrow host specificity requiring precise diagnosis, and the need for robust regulatory frameworks. The future lies in synthetic biology, engineering phages to enhance lytic efficiency, expand host range, and deliver biofilm-degrading enzymes directly to the infection niche.
Skin Microbiome
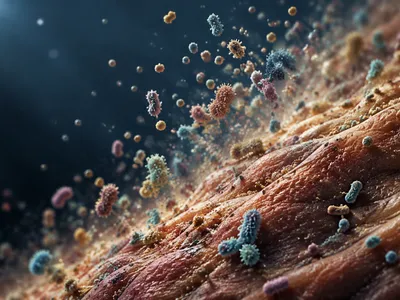
The skin, our largest organ, hosts a diverse and site-specific microbiota that is integral to cutaneous homeostasis and overall immune defense. This ecosystem varies remarkably between dry, moist, and sebaceous regions, with genera like Cutibacterium, Staphylococcus, and Malassezia being predominant.
A balanced skin microbiome prevents colonization by pathogens through competitive exclusion and the production of antimicrobial peptides (AMPs). Disruption of this balance, or dysbiosis, is a key factor in the pathogenesis of conditions such as atopic dermatitis, psoriasis, and acne vulgaris.
Advanced genomic sequencing has elucidated the functional capacity of the skin microbiome beyond mere taxonomy. Commensals like Staphylococcus epidermidis contribute to the acidic pH barrier by fermenting sebum triglycerides into free fatty acids. Furthermore, they engage in a constant dialog with keratinocytes and resident immune cells, modulating the local inflammatory tone. For instance, certain microbial byproducts can activate toll-like receptors (TLRs) on keratinocytes, leading to the controlled production of defensins and cytokines that prime the skin's defenses without eliciting overt inflammation. This sophsticated, localized immunomodulation underscores the skin microbiome's role as an active, dynamic component of the integumentary system's defense architecture.
Environmental Metagenomics Unlocking Unseen Worlds
Environmental metagenomics, the direct genetic analysis of microbial communities from their natural habitats, has revolutionized our understanding of planetary biodiversity and biogeochemical cycles. This culture-independent approach bypasses the limitations of traditional microbiology.
By sequencing the collective DNA from soil, ocean, or air samples, scientists have uncovered millions of previously unknown genes and metabolic pathways. This "genomic dark matter" holds immense potential for biotechnology and medicine.
Discoveries include novel extremophilic enzymes from hydrothermal vents useful in industrial processes and previously uncultivated bacteria from soil that produce new classes of antibiotics, addressing the antibiotic discovery pipeline crisis.
This field has also critically informed public health through pathogen surveillance and antimicrobial resistance tracking. By analyzing wastewater and urban microbiomes, researchers can map the prevalence and spread of antibiotic resistance genes (ARGs) in near real-time, providing an early warning system for emerging health threats and enabling more targeted interventions to curb the spread of superbugs in communities.
Microbial Communication Quorum Sensing
Bacteria coordinate collective behaviors through a cell-cell communication process called quorum sensing (QS), which relies on the production, release, and detection of small diffusible signaling molecules known as autoinducers. This sociomicrobiological mechanism allows a population to sense its density and synchronize gene expression accordingly.
QS regulates phenotypes critical for virulence, such as biofilm formation, toxin production, and bioluminescence. Targeting QS, known as quorum quenching, represents a promising anti-virulence strategy that disarms pathogens without imposing lethal selective pressure, potentially reducing the development of resistance.
The therapeutic potential of quorum quenching is being explored using enzymatic degradation of autoinducers, synthetic antagonist molecules, and competitive inhibition. These approaches aim to disrupt pathogenic coordination and enhance the efficacy of conventional antimicrobials, especially in treating resilient biofilm-associated infections like those in cystic fibrosis or on medical implants.
| QS System Type | Key Signaling Molecule | Example Bacterium & Regulated Function |
|---|---|---|
| LuxI/LuxR-type (AHL-based) | N-acyl homoserine lactones (AHLs) | Pseudomonas aeruginosa (biofilm, virulence factors) |
| AI-2 System | Autoinducer-2 (furanosyl borate diester) | Vibrio harveyi (bioluminescence); considered a "universal" signal |
| Oligopeptide-based (Gram+) | Modified oligopeptides | Staphylococcus aureus (toxin production, agr system) |
Beyond single-species systems, interspecies and interkingdom signaling reveal a staggering complexity in microbial ecosystems. For instance, gut commensals may utilize QS molecules to influence host epithelial cell responses, while plants and animals have evolved receptors to detect bacterial autoinducers, modulating their own immune defnses. This intricate cross-talk underscores that microbial communities function as highly integrated, information-processing networks, and future medical interventions may increasingly focus on managing this communication rather than merely killing the microbes, paving the way for next-generation probiotics and targeted microbiome modulators.
Probiotics and Postbiotics From Concept to Counter
The clinical application of beneficial microbes has evolved from the traditional concept of probiotics—live microorganisms that confer a health benefit—to include postbiotics, which are preparations of inanimate microorganisms and/or their components that provide a physiological benefit. This shift addresses challenges related to probiotic viability, storage, and safety in immunocompromised hosts.
Postbiotics include metabolites, cell wall fragments (e.g., peptidoglycan, teichoic acids), and secreted proteins. These components can directly modulate host pathways, offering more consistent and targeted effects than live bacteria, whose activity is variable and dependent on colonization success.
Evidence supports the use of specific probiotic strains, such as Lactobacillus rhamnosus GG and Saccharomyces boulardii, for managing antibiotic-associated diarrhea and acute gastroenteritis. The mechanisms include competitive exclusion of pathogens, enhancement of gut barrier function, and modulation of the immune response.
For postbiotics, research highlights the efficacy of bacterial lysates and short-chain fatty acid preparations. Butyrate supplements, for example, are investigated for their anti-inflammatory effects in ulcerative colitis, while specific cell-free supernatants from lactobacilli show promise in alleviating symptoms of irritable bowel syndrome by influencing visceral hypersensitivity and motility.
- Strain-Specificity: Effects are highly dependent on the precise microbial strain, not just the species or genus.
- Dose-Response Relationship: Clinical outcomes are directly linked to the administered dose and formulation.
- Ecological Context: Efficacy depends on the recipient's existing gut microbiota composition and metabolic state.
- Defined Mechanisms: Modern research focuses on identifying specific molecular actors (e.g., surface proteins, metabolites) responsible for the health effect.
The future of this field lies in precision microbiome interventions, moving beyond generic supplements to personalized formulations based on an individual's microbiomic and metabolomic profile. Engineered probiotics and synthetically produced postbiotics, designed to deliver specific therapeutic molecules (e.g., cytokines, enzymes) directly to the gut environment, represent the cutting edge. Furthermore, regulatory frameworks are adapting to classify these novel products appropriately, ensuring their safety and efficacy through rigorous, mechanism-based clinical trials that validate health claims beyond traditional nutritional supplement standards.